Export modules
Export modules allow to export computed data into external software for
post-processing. The number of export module records is specified in
analysis record using nmodules parameter (see the initial part of section 3). The general format
is the following:
| *EntType |
[tstep_all] |
|
|
[tstep_step #(in)] |
|
|
[tsteps_out #(rl)] |
|
|
[subtsteps_out #(in)] |
|
|
[domain_all] |
|
|
[domain_mask #(in)] |
To select all solution steps, in which output will be performed, use
tstep_all. To select each tstep_step-nth step, use
tstep_step parameter. In order to select only specific
solution steps, the tsteps_out list can be specified,
supplying solution step number list in which output will be done.
To select output for all domain of the problem the domain_all
keyword can be used. To select only specific domains,
domain_mask array can be used, where the values of the array
specify the domain numbers to be exported.
If the parameter subtsteps_out = 1, it turns on the export of intermediate results,
for example during the substepping or individual equilibrium iterations. This option requires support from the solver.
Currently, the supported export modules are following
- VTK export, DEPRECATED - Use VTKXML
| vtk |
[vars #(ia)] |
| |
[primvars #(ia)] |
| |
[cellvars #(ia)] |
| |
[stype #(in)] |
| |
[regionstoskip #(ia)] |
| vtkxml |
[vars #(ia)] |
| |
[primvars #(ia)] |
| |
[cellvars #(ia)] |
| |
[ipvars #(ia)] |
| |
[stype #(in)] |
| |
[regionsets #(ia)] |
| |
[timeScale #(rn)] |
- The vtk module is obsolete, use vtkxml instead. Vtkxml allows to export results recovered on region by region basis and has more features.
- The array vars contains identifiers for those internal
variables which are to be exported. These variables will be smoothed and transfered to nodes. The id values are defined by
InternalStateType enumeration, which is defined in include file
``src/oofemlib/internalstatetype.h''.
- The array primvars
contains identifiers of primary variables to be exported.
The possible values correspond to the values of enumerated type
UnknownType, which is again defined in ``src/oofemlib/unknowntype.h''.
Please note, that the values corresponding to enumerated type values
start from zero, if not specified directly and that not all values are
supported by particular material model or analysis type.
- The array cellvars contains identifiers of constant variables defined
on an element (cell), e.g. a material number. Identifier numbers are specified in
``src/oofemlib/internalstatetype.h''.
- The array ipvars contains identifiers for those internal
variables which are to be exported. These variables will be directly exported (no smoothing) as point dataset, where each point corresponds to individual integration point. A separate vtu file for these raw, point data will be created.
The id values are defined by
InternalStateType enumeration, which is defined in include file
``src/oofemlib/internalstatetype.h''.
- The parameter stype allows to select smoothing procedure for
internal variables, which is used to compute nodal values from values
in integration points. The supported values are 0 for simple nodal averageing (generally supported only by triangular and tetrahedral elements),
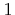 for Zienkiewicz Zhu recovery (default), and
for Zienkiewicz Zhu recovery (default), and 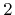 for Superconvergent Patch Recovery (SPR, based on least square fitting).
for Superconvergent Patch Recovery (SPR, based on least square fitting).
- The export is done on region basis, on each region, the nodal recovery is performed independently and results are exported in a separate piece. This allows to take into account for discntinuities, or to export variables defined only by particular material model. The region volumes are defined using sets containing individual elements. By default the one region is created, containing all element in the problem domain. The optional parameter regionsets allows to use user-defined. The individual values refer to numbers (ids) of domain sets. Note, that regions are determined solely using elements.
- timeScale scales time in output. In transport problem, basic units are seconds. Setting timeScale = 2.777777e-4 (=1/3600.) converts all time data in vtkXML from seconds to hours.
By default vtk and vtkxml modules perform recovery over the whole domain. The VTKXML module can operate in region-by-region mode (see nvr and vrmap parameters). In this case, the smoothing is performed only over particular virtual region, where only elements in this virtual region participate.
- Homogenization of IP quantities in the global coordinate system (such as stress, strain, damage, heat flow). Corresponding IP quantities are summed and averaged over the volume. It is possible to select region sets from which the averaging occurs. The averaging works for all domains with an extension to trusses. A truss is considered as a volume element with oriented stress and strain components along the truss axis. The transformation to global components occurs before averaging.
| |
ists #(ia) |
| |
[scale #(rn)] |
| |
[regionSets #(ia)] |
- An integer array ists specifies internal state types for export which are defined in internalstatetype.h file.
- The parameter scale multiplies all averaged IP quantities. scale=1 by default.
- An integer array regionSets specifies region sets for averaging. All domain is averaged by default.
- Gauss point export is useful if one needs to plot a certain variable
(such as damage) as a function of a spatial coordinate using tools like
gnuplot. It generates files with data organized in columns, each row
representing one Gauss point. In this way, one can plot e.g. the damage
distribution along a one-dimensional bar.
| gpexportmodule |
[vars #(ia)] |
| |
[ncoords #(in)] |
- The array vars contains identifiers for those internal
variables which are to be exported. The id values are defined by
InternalStateType enumeration, which is defined in include file
``src/oofemlib/internalstatetype.h''.
- Parameter ncoords specifies the number of spatial coordinates to be exported at each Gauss point. Depending on the spatial dimension of the
domain, the points can have one, two or three coordinates. If ncoords
is set to -1, only those coordinates that are actually used are exported.
If ncoords is set to 0, no coordinates are exported.
If ncoords
is set to a positive integer, exactly ncoords coordinates are exported.
If ncoords exceeds the actual number of coordinates, the actual
coordinates are supplemented by zeros. For instance, if we deal with a 2D
problem, the actual number of coordinates is 2. For ncoords=3,
the two actual coordinates followed by 0 will be exported.
For ncoords=1, only the first coordinate will be exported.
The Gauss point export module creates a file with extension ``gp''
after each step for which the output is performed. This file contains
a header with lines starting by the symbol #, followed by the actual data
section.
Each data line corresponds to one Gauss point and contains the following
data:
- element number,
- material number,
- Gauss point number,
- contributing volume around Gauss point,
- Gauss point global coordinates (written as a real array of length ncoords),
- internal variables according to the specification in vars (each written as a real array of the corresponding length).
Example:
``GPExportModule 1 tstep_step 100 domain_all ncoords 2 vars 5 4 13 31 64 65''
means that the *.gp file will be written after each 100 steps and will contain
for each of the Gauss points in the entire domain its 2 coordinates and also internal
variables of type 4, 13, 31, 64 and 65, which are the strain tensor, damage tensor, maximum equivalent strain level, stress work density and dissipated work density. Of course, the material model must be able to deliver such variables.
The size of the strain tensor depends on the spatial dimension, and the size
of the damage tensor depends on the spatial dimension and type of model
(e.g., for a simple isotropic damage model it will have just 1 component
while for an anisotropic damage model it may have more). The other variables
in this example are scalars, but they will be written as arrays of length 1,
so the actual value will always be preceded by ``1'' as the length of the array. Since certain internal variables have the meaning of densities (per unit volume
or area, again depending on the spatial dimension), it is useful to have
access to the contributing volume of the Gauss point. The product of this
contributing volume and the density gives an additive contribution to the
total value of the corresponding variable. This can be exploited e.g. to
evaluate the total dissipated energy over the entire domain.
Borek Patzak
2018-01-02